I am giving a talk about using R for survival analysis and I wanted to talk first about the Kaplan-Meier curve and how you might draw it in R.
I wrote about the Kaplan-Meier curve in a previous webpage - but that was a generic example. First - you read the data in R and transform it into a survival object.
day <- c(37,40,43,44,45,
47,49,54,56,58,59,60,61,62,68,
NA,71,NA,NA,75,
NA,NA,89,NA,96)
death <- is.finite(day)
tim <- day
tim[is.na(day)] <- 70
library("survival")
fly.surv <- Surv(tim,death)The Surv function has options for left censored and interval censored observations. Read the help file for details.
There is a print method for survival objects.
print(fly.surv)The survival object prints with a “+” attached to any censored observation.
[1] 37 40 43 44 45 47 49 54 56 58 59 60 61 62 68 70+ 71 70+ 70+ 75 70+ 70+ 89
[24] 70+ 96The survfit function creates a new object that summarizes the data in a survival object using a Kaplan-Meier curve or a Cox regression model. The input for survfit is a formula with a survival object on the left side of the equation. A model with “~1<U+2033> fits a single Kaplan-Meier curve to the entire survival object.
fly.fit <- survfit(fly.surv~1)There is a print method for survfit objects .
print(fly.fit)which lists some basic statistics
Call: survfit(formula = fly.surv ~ 1)
records n.max n.start events median 0.95LCL 0.95UCL
25 25 25 19 61 56 NAThere is also a summary method
summary(fly.fit)which produces a more detailed set of statistics.
Call: survfit(formula = fly.surv ~ 1)
records n.max n.start events median 0.95LCL 0.95UCL
25 25 25 19 61 56 NA
time n.risk n.event survival std.err lower 95% CI upper 95% CI
37 25 1 0.96 0.0392 0.8862 1.000
40 24 1 0.92 0.0543 0.8196 1.000
43 23 1 0.88 0.0650 0.7614 1.000
44 22 1 0.84 0.0733 0.7079 0.997
45 21 1 0.80 0.0800 0.6576 0.973
47 20 1 0.76 0.0854 0.6097 0.947
49 19 1 0.72 0.0898 0.5639 0.919
54 18 1 0.68 0.0933 0.5197 0.890
56 17 1 0.64 0.0960 0.4770 0.859
58 16 1 0.60 0.0980 0.4357 0.826
59 15 1 0.56 0.0993 0.3956 0.793
60 14 1 0.52 0.0999 0.3568 0.758
61 13 1 0.48 0.0999 0.3192 0.722
62 12 1 0.44 0.0993 0.2827 0.685
68 11 1 0.40 0.0980 0.2475 0.646
71 4 1 0.30 0.1136 0.1428 0.630
75 3 1 0.20 0.1114 0.0672 0.596
89 2 1 0.10 0.0900 0.0171 0.584
96 1 1 0.00 NaN NA NAMost importantly - there is a plot method.
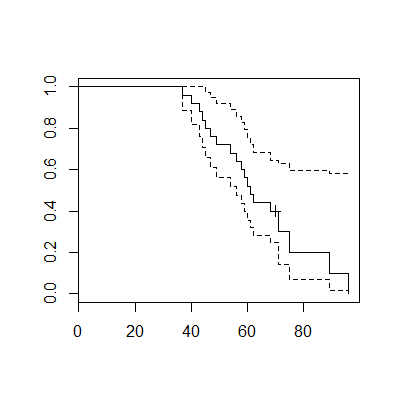
plot(fly.fit)The graph includes the survival curve (either from a Kaplan-Meier estimate or a Cox regression model) and confidence limits. The graph displays a “+” at any censored value.
There’s a lot more on survival models which I hope to cover in another blog entry.